Approaches
In our lab, we combine methods from multiple disciplines according to the question at hand. Here we introduce our research topics and the techniques we use through the lens of the fields they draw on.
Evolutionary Biology
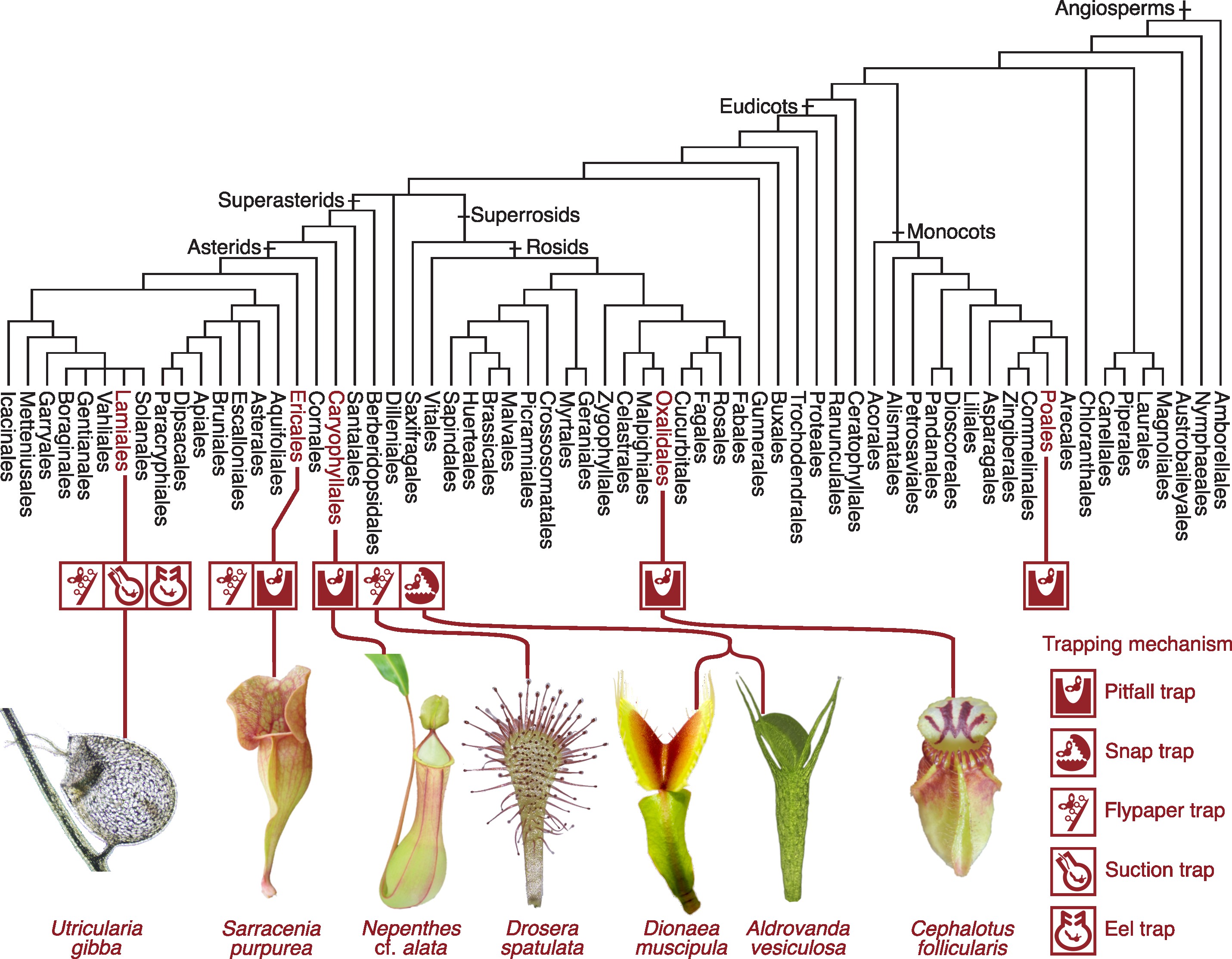
Just as children differ slightly from their parents, organisms also change gradually from generation to generation. If we stretch that timescale far enough, we arrive at the familiar story that primitive bacteria gave rise, over roughly four billion years, to humans, frogs, and carnivorous plants. That story is correct, but it leaves many questions open: what mechanisms drove those transformations, and what key events shaped major evolutionary transitions? Our lab works on those questions.
Plant Molecular Biology
Unlike animals, plants cannot easily move to a better place when conditions become too hot, too cold, or too poor in resources. Instead, they survive by changing their own bodies and physiology in response to the environment. We study that way of life in plants.
Molecular biology, centered on nucleic acids and proteins, is a core foundation of modern biology and one of the main pillars of our lab. Plant experiments require workflows tailored to plant cells, including the removal of secondary metabolites during nucleic-acid extraction and the use of Agrobacterium for gene delivery. Agrobacterium is also commonly used in CRISPR-based gene editing. We are developing genetic manipulation techniques for carnivorous plants as well.
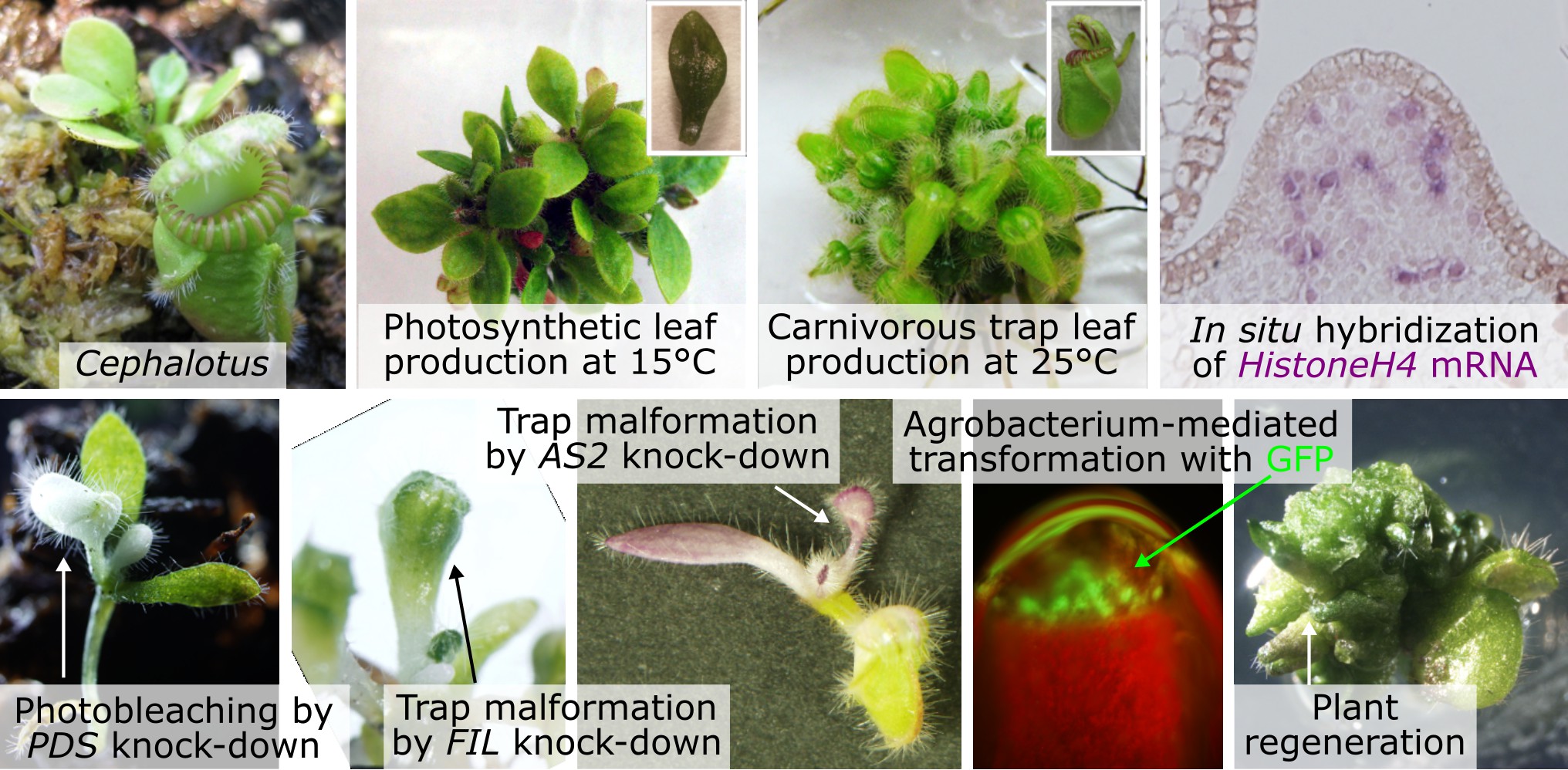

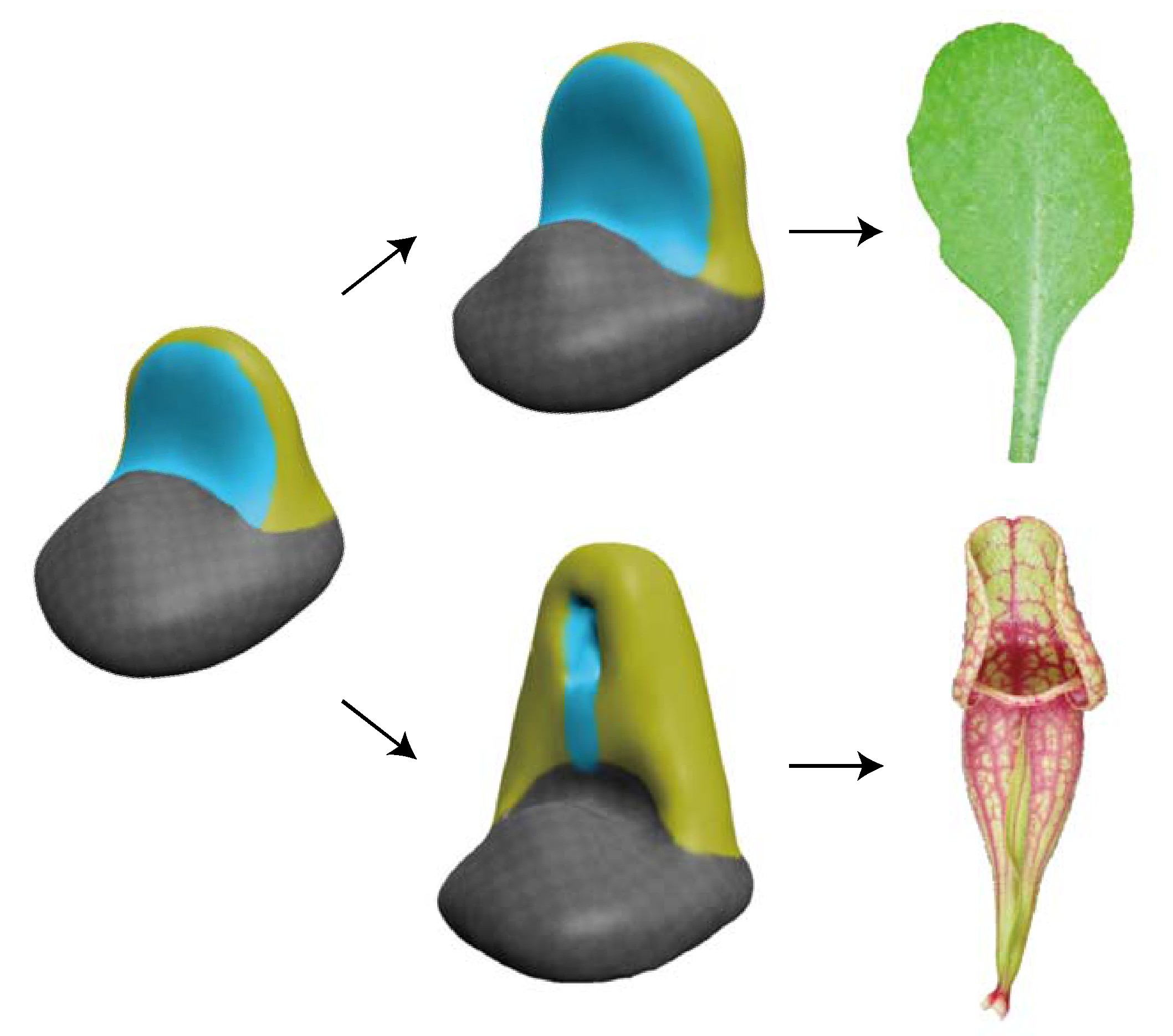
Evo-Devo
Carnivorous trapping leaves have highly complex forms, but how were they acquired through evolution? It is difficult to infer the path from ordinary leaves to mature traps by looking only at the final structures. Much of that complexity, however, is built step by step during development. Early in development, ordinary leaves and trapping leaves are simple and remarkably similar. By carefully tracing when and how they diverge, we can infer the evolutionary events that were important for the origin of trapping-leaf morphology.
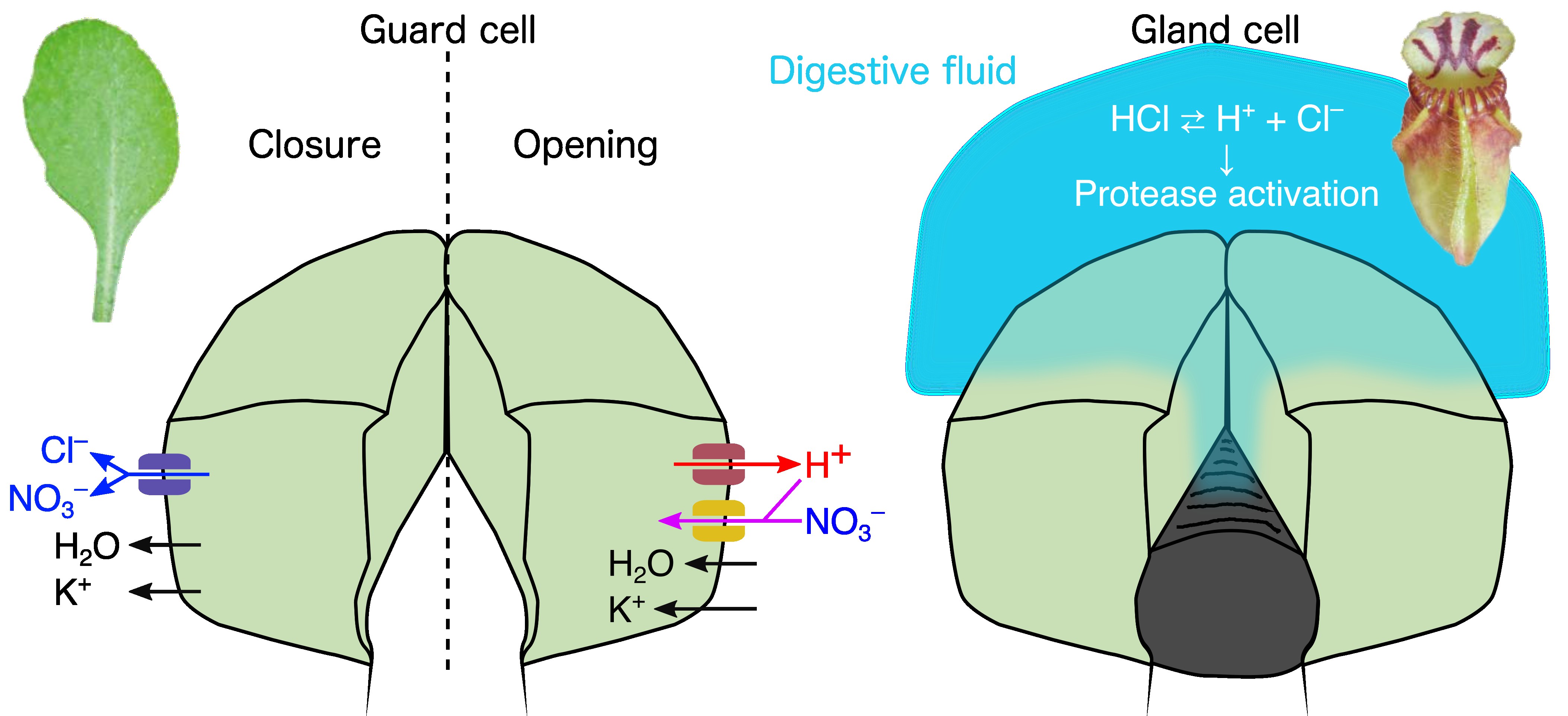
Plant Physiology and Electrophysiology
Appearance is only part of the story. To understand the digestive capacity and nutrient-absorption mechanisms of carnivorous plants, we also need to study transport, metabolism, and plant hormones, all of which belong to plant physiology.
Electricity is also often important. When people hear “bioelectricity,” they tend to think of muscles and nerves, but plants use electrical signals too. These signals are based on membrane potential, the voltage difference created by unequal ion composition across biological membranes. Electrophysiological methods for detecting membrane potentials are useful not only for analyzing movements such as the snap of the Venus flytrap, but also for studying more general carnivorous-plant functions, including digestion and absorption, where ion transport plays a major role. Kenji does not currently practice these techniques himself, but if experienced members join the lab we would like to build in-house capability. For now, we carry out the necessary electrophysiological analyses through collaboration.
Biochemistry and Protein Engineering
Proteins can be synthesized and modified relatively easily in vitro. With careful experimental design, this makes it possible to obtain high-resolution insights into biological evolution. Biochemical and protein-engineering approaches are therefore powerful tools in our research.
Synthetic Biology
The “build to understand” approach is highly effective in science. If we identify key principles from the study of carnivorous plants, one way to test them is to partially recreate carnivorous traits in ordinary plants. Re-engineering Arabidopsis thaliana into a full Nepenthes-like carnivore is not yet realistic, but introducing a single carnivorous-plant gene to alter some aspect of nutrient metabolism, or inducing hair-like structures reminiscent of those found in carnivorous plants, is becoming increasingly plausible.
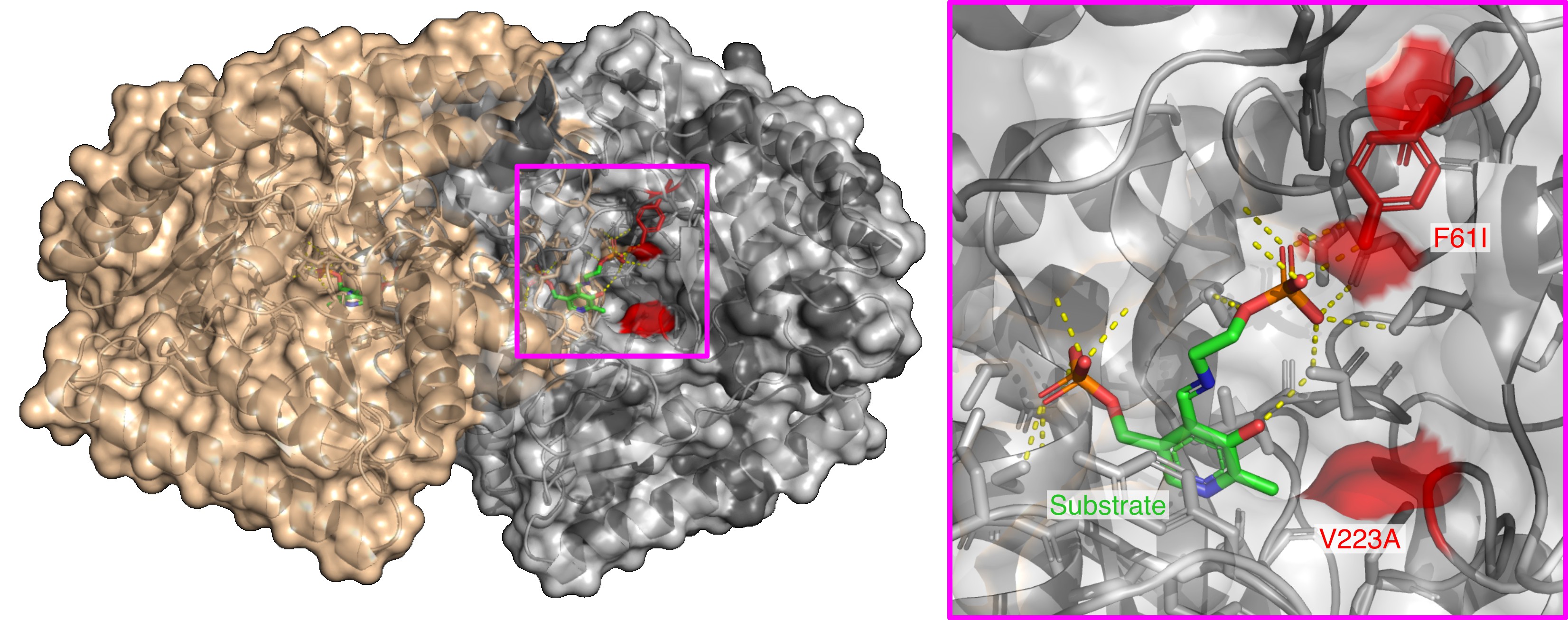
At the protein level, the idea is even more tractable: we can introduce mutations and ask whether they confer characteristic properties of carnivorous-plant proteins. Starting from reconstructed ancestral proteins, this approach becomes a kind of experimental replay of evolution. Comparing the mutations that actually occurred with plausible alternatives can also tell us something about evolutionary paths that were not taken. Broadly speaking, these are synthetic-biology-like approaches.
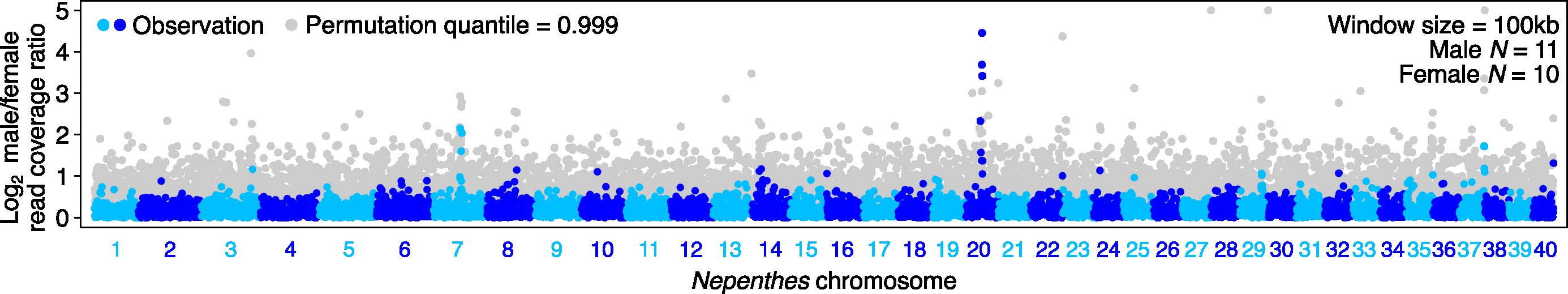
Genomics
The vast amount of information encoded in genomes provides crucial clues to the origin and biology of organisms. Producing whole-genome sequences, predicting gene structures, and inferring gene function are important groundwork for making biological discoveries from genomic data. By combining long-read sequencing platforms such as PacBio and Oxford Nanopore with Hi-C, we can obtain chromosome-scale genome assemblies. Generating the sequence data requires molecular biology, and much of the downstream analysis relies on bioinformatics.

Bioinformatics
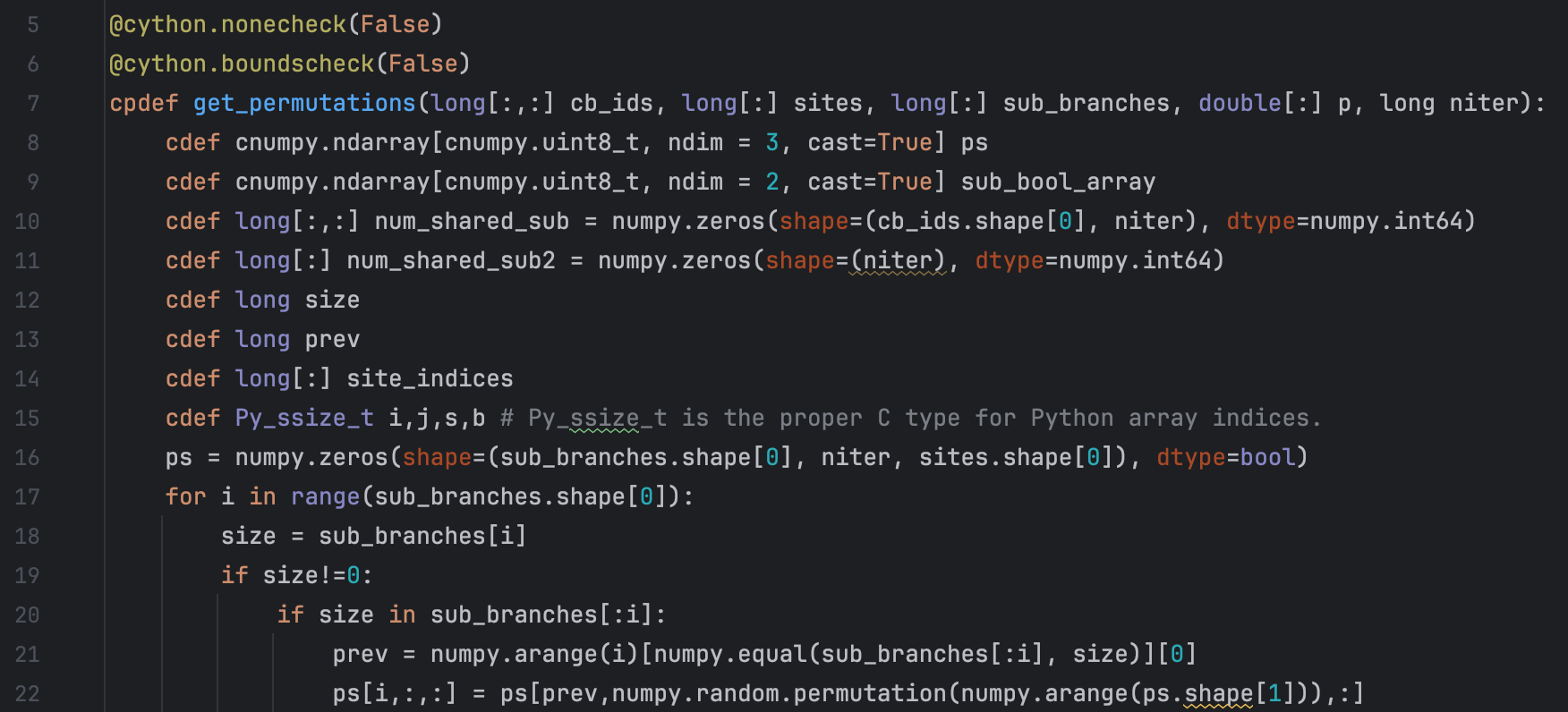
When we confront complex biological phenomena, bioinformatics is an extremely powerful tool. Using existing tools for omics and comparative genomics already allows us to extract a great deal of insight, but our lab also develops new methods.
Phylogenetics
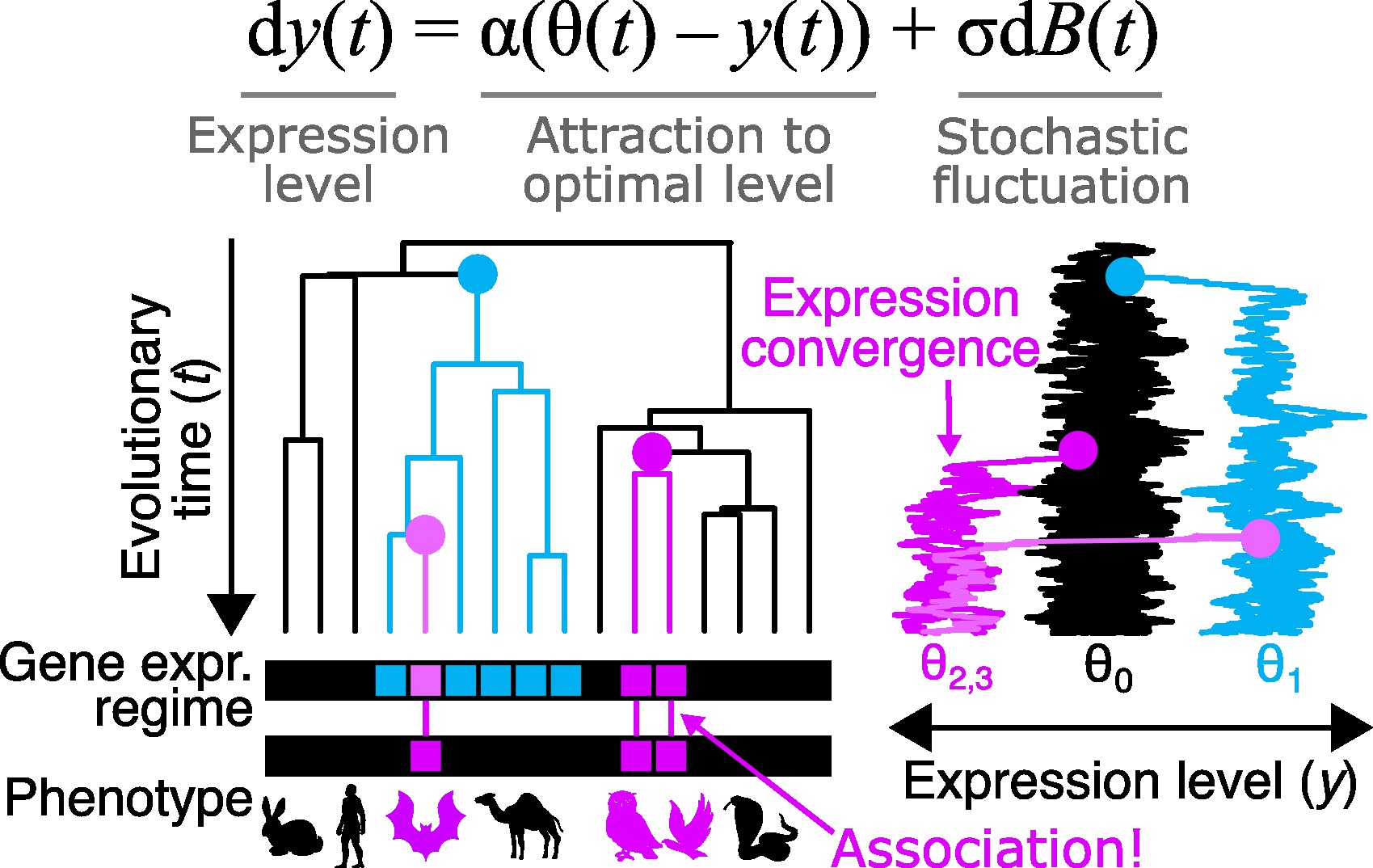
Phylogenetics is a classical area within bioinformatics, but it remains a core framework for comparative genomics. In addition to inferring evolutionary relationships among organisms, we routinely use phylogenetic comparative methods to model the evolution of gene expression levels and other traits on phylogenetic trees.
Statistics
Statistics is an essential skill in modern biology. Through day-to-day research, our lab provides opportunities to learn statistical thinking from the basics to more advanced applications, including how to describe statistical analyses clearly in papers.